Leaving Community
Are you sure you want to leave this community? Leaving the community will revoke any permissions you have been granted in this community.
For each dataset, you can submit predictions for protein-ligand poses and/or for protein-ligand affinity scores or rankings. All predictions must be submitted in the form of gzipped tar (.tgz) files of two possible types:
- A Dock tgz file containing pose predictions, a protocol file explaining how the predictions were made, and a user-information file. Optionally, it may also contain scores files with structure-based protein-ligand scores and corresponding rankings based on the predicted poses, and corresponding scoring protocol files explaining how these scores were computed. Each pose must be represented by two separate files, in different file formats: a PDB file for the protein, and an MDL mol file (see, e.g., http://en.wikipedia.org/wiki/Chemical_table_file) for the ligand. Any ligand coordinates provided in PDB format or included in the protein PDB files will be ignored. (We are asking for molfiles to prevent problems with the parsing of ligand coordinates in PDB format, which arose in Grand Challenge 2015.)
- A Score tgz file containing protein-ligand scores and corresponding rankings, without any pose predictions, and is to be used for both purely ligand-based prediction methods, such as QSAR; and structure-based predictions when there are no crystallographic poses to compare with. It also must contain a protocol file for each set of scores and ranks, explaining how they were computed, as well as a user-information file.
Note that challenge PL-2016-1 is unusual because it includes cases where the common entity is the ligand, and it is the proteins that must be scored. In particular, you are challenged to score:
- 17-OHP: proteins PL-2016-1-O-1 and PL-2016-1-O-2 against ligand 17-OHP
- Cholecalciferol:
- proteins PL-2016-1-C-1, PL-2016-1-C-2, and PL-2016-1-C-3, against 25-D3
- ligands 25-D3 and vitD3 against protein PL-2016-1-C-1
For the sake of clarity, the scores files for items 1 and 2a will be termed ProteinScores files, while those for item 2b will be termed LigandScores files.
The structures of the dock and score tgz files for this challenge are summarized in the figure below, and the subsequent text details their contents and format. Additionally, blank template files for both components of this challenge are available for download, as are examples of completed Dock.tgz and Score.tgz files. Note that the example files contain artificial information, and thus serve only to illustrate the required contents and formats of a submission. Please adhere to the structure of the template files. Files that do not adhere to the requirements may be rejected by our submission system. In addition, it's possible that files are accepted to the system and yet do not follow the format requirements. In these instances, we do our best to accommodate the users and interpret the submission, but in some instances, we may find the file too difficult to work with, and in those cases, we reserve the right to omit the submission from evaluation.
Example and Template Files
Cholecalciferol: cholecalciferol-examples-and-templates.tgz
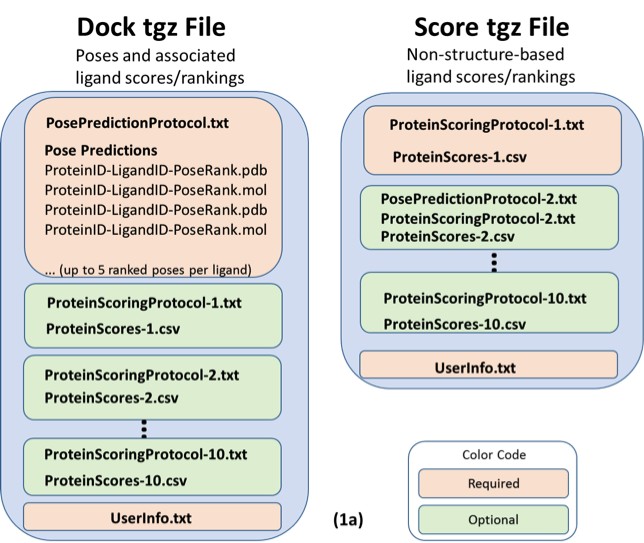
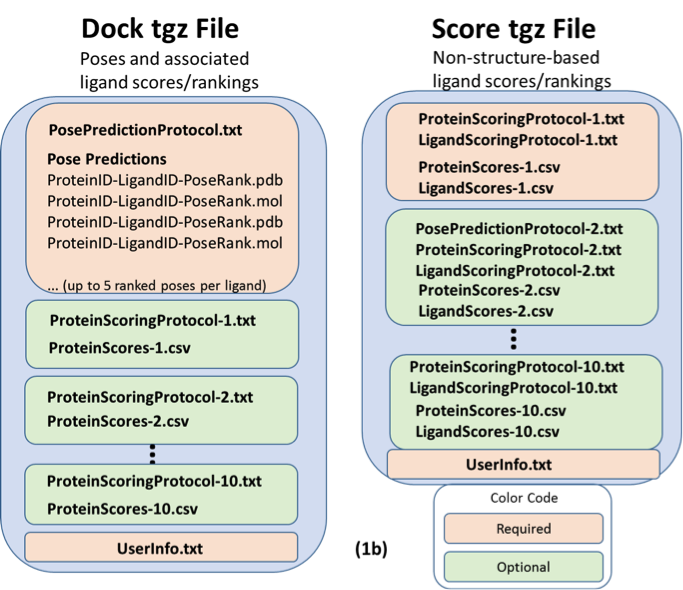
Figure 1. Diagrammed contents of the two types of *tgz files for both the 17-OHP (a), and cholecalciferol (b), components of the PL-2016-1 Challenge. 17-OHP predictions should contain ProteinScore files only. Cholecalciferol predictions should include both ProteinScore and LigandScore files. Note that the 17-OHP Score file example (1a, right) includes a first set of scores that are not structure-based, so no pose prediction protocol is included (prediction set 1), followed by a second set of scores which is structure-based, so a pose prediction protocol file is included (prediction set 2).
Dock tgz file
A Dock tgz file is used to submit one set of structure-based predicted protein-ligand poses for challenge cases where crystallographic structures are available, where up to five poses are permitted for each protein-ligand pair. The file must contain a description of and the results from one and only one pose-prediction protocol. Optionally, the file may also contain the protocols for and results from up to ten different protein-ligand affinity scoring and ranking calculations, where the scores must be based on the poses in the same dock tgz file. If more than one scoring protocol and results set is included in the file, the files describing them must have distinct names. The name of a dock tgz file must include the string "dock", so that the file name has the form *dock*.tgz, except that the string "score" may not be in the file name (see below).
The following subsections detail the files contained in a Dock .tgz file.
Pose prediction protocol and result files
One Dock tgz file contains a single docking protocol file, and up to 5 protein structure PDB files and 5 ligand mol file poses predicted by this protocol for each ligand. The protocol file, named PosePredictionProtocol.txt, contains a brief, structured summary, in the form of a plain-text document, of how you did the pose predictions. Lines beginning with a hash-tag (#) may be included as comments. The file must contain the following components, as illustrated in the template and example:
- Your informal, brief name for the protocol
- A list of the major software packages and their versions used for system preparation and pose prediction
- List of key parameters used for system preparation
- Plain-text description of system preparation method
- List of key parameters used for pose prediction
- Plain-text description of pose prediction method
Each item must begin with the appropriate keyword; respectively:
- Name:
- Software:
- System Preparation Parameters:
- System Preparation Method:
- Pose Prediction Parameters:
- Pose Prediction Method:
Each pose prediction must be provided in the form of a protein structure PDB file, and a corresponding ligand mol file with 3D atomic coordinates for the pose, where the coordinates in the protein PDB file and the ligand molfile are in the same frame of reference. Note that you may treat the protein as rigid or flexible, however, you must rotationally and translationally superimpose all of your final structure predictions, for both the 17-OHP and cholecalciferol cases, onto structure PL-2016-1-O-1, to facilitate evaluation of your predictions.
The file names of your pose prediction protein PDB and ligand mol files must be constructed as follows:
<PDB ID of initial protein structure>-<LigandID>-<poseRank>.pdb
<PDB ID of initial protein structure>-<LigandID>-<poseRank>.mol
Here
PL-2016-1-C-1-25-D3-2.mol
PL-2016-1-C-1-25-D3-2.pdb
Additionally, if you submit multiple poses for a ligand, then the first line of each molfile must take the form
REMARK
For example, this line might be
REMARK energy -20.6
REMARK score 5.7
Energies must be in kcal/mol; scores may be in arbitrary units.
Sample pose prediction protein PDB files and ligand mol files are included with these instructions, as are example and template files for PosePredictionProtocol.txt.
Protein/ligand scoring protocol and result files
Each protein/ligand scoring and ranking is described by two files: a protein or ligand scoring protocol file, named either ProteinScoringProtocol-n.txt or LigandScoringProtocol-n.txt, respectively, where n is n integer from 1 to a maximum of 10; and a protein or ligand scoring results file, named ProteinScores-n.csv or LigandScores-n.csv, respectively, which contains the scores and corresponding rankings generated by the corresponding protocol.
The protein/ligand scoring protocol file must contain a brief, structured summary, in the form of a plain-text document, of how you scored the proteins or ligands according to predicted affinity for the target protein. A template file is provided for your convenience, as is a sample filled-out file. Lines beginning with a hash-tag (#) may be included as comments. The file must contain the following components, as illustrated in the template and example:
- Your informal brief name for the protocol
- A list of the major software packages and their versions used in the protocol
- A listing of the key parameters used in the calculations
- A brief narrative of the procedure.
Each item must begin with the appropriate keyword; respectively:
- Name:
- Software:
- Parameters:
- Method:
The protein/ligand scoring results file lists your rankings and scores or energies of the binding strengths of the ligands to the respective proteins. Again, lines beginning with # will be treated as comments. Since some scoring methods provide results interpretable as binding energies or free energies, while others provide scores without well-defined units, the first non-comment line of your file must state whether you are providing energies or scores. This line must take one of the following forms:
Type: energy
orType: score
If your results are given as energies, the units must be in units of kcal/mol.
Each subsequent non-comment line of the file comprises the identifier of one protein/ligand for the protein target or ligand in question; your ranking of the protein or ligand within the set, where 1 corresponds to maximal affinity; and your computed binding energy, free energy, or score. These three items should be separated by commas.
The scoring file must contain a line for every ligand or protein in the challenge. If you have not entered a prediction for a ligand or protein, the corresponding line should have a placeholder: "inact" for compounds you identified as inactive, or "nopred" if you are not supplying a prediction for the compound for any reason.
Please refer to the template and example files; note that the template files are prefilled with the list of protein or ligand identifiers, for your convenience.
User Information File
The user-information file is a text file named UserInfo.txt and containing five lines of text, as follows:
- Submitter Last Name:
- Submitter First Name:
- Submitter email:
- Submitter Organization:
- Research group or PI Name:
- Research group or PI Email:
We are asking for this file to maximize clarity regarding the associations between submissions and submitters and research groups.
Score tgz file
A Score tgz file is used to submit one to ten sets of protein/ligand scores or rankings, which have been generated by a ligand-based method, such as QSAR; or by a structure-based method, in cases where crystallographic structures are not available for comparison. If more than one scoring protocol and results set is included in the file, the files describing them must have distinct names. The name of a Score tgz file must include the string "score", so that the file name has the form *score*.tgz, except that the string "dock" may not be in the file name.
If a prediction set is structure-based, you must provide a pose-prediction protocol, a scoring protocol based on the predicted poses, and a set of scores, based on the instructions given above for the Dock tgz file. However, actual poses should not be submitted, as there are no crystallographic poses available for reference. If a prediction set is ligand-based, there are no poses, so you must provide only a scoring protocol and a set of scores, again based on the instructions given above for the Dock tgz file. In either case, you must also include the user information file. Each set of files should be given its own integer index. For example, if you submit ligand-based results followed by structure based results, you might submit the following files:
ProteinScoringProtocol-1.txt
ProteinScores-1.csv
PosePredictionProtocol-2.txt
ProteinScoringProtocol-2.txt
ProteinScores-2.csv
How to make your Dock and Score tar files
In order to enable automated processing of all submissions, we ask that you generate your tar files as follows, for both the Dock and Score tgz files.
- Put all files in one directory, with any name of your choosing.
- There should be no pathnames beyond that of the single directory. For example, your directory should not be named "/home/username/cholecalciferol/Dock". The following is an example tar command which will remove the extra directories:
tar -cvzf myDock.tar.gz --directory= /home/username/cholecalciferol Dock - There should be no extraneous files, such as .sh scripts, Excel files, other tar files, etc.
- For Mac users, be sure your tar file includes no extraneous Mac-specific files. For example, these might show up as "._ PL-2016-1-C-1-25-D3-1.pdb" or ".DS_Store". The following example, tar command will generate a tar file without these extras:
tar --disable-copyfile --exclude=.DS_Store -cvzf myDock.tar.gz Dock - Each submission must have a unique filename.
- Example "Dock" file output from "tar -tzf" command. Notice the "top level" Dock directory.
Dock/
Dock/PL-2016-1-O-1-17-OHP-1.mol
Dock/PL-2016-1-O-1-17-OHP-1.pdb
Dock/PL-2016-1-O-1-17-OHP-2.mol
Dock/PL-2016-1-O-1-17-OHP-2.pdb
Dock/PL-2016-1-O-1-17-OHP-3.mol
Dock/PL-2016-1-O-1-17-OHP-3.pdb
Dock/PL-2016-1-O-1-17-OHP-4.mol
Dock/PL-2016-1-O-1-17-OHP-4.pdb
Dock/PL-2016-1-O-1-17-OHP-5.mol
Dock/PL-2016-1-O-1-17-OHP-5.pdb
Dock/PL-2016-1-O-2-17-OHP-1.mol
Dock/PL-2016-1-O-2-17-OHP-1.pdb
Dock/PL-2016-1-O-2-17-OHP-2.mol
Dock/PL-2016-1-O-2-17-OHP-2.pdb
Dock/PL-2016-1-O-2-17-OHP-3.mol
Dock/PL-2016-1-O-2-17-OHP-3.pdb
Dock/PL-2016-1-O-2-17-OHP-4.mol
Dock/PL-2016-1-O-2-17-OHP-4.pdb
Dock/PL-2016-1-O-2-17-OHP-5.mol
Dock/PL-2016-1-O-2-17-OHP-5.pdb
Dock/PosePredictionProtocol.txt
Dock/ProteinScores-1.csv
Dock/ProteinScoringProtocol-1.txt
Dock/UserInfo.txt

